Resources
Lectures
Decision Making in Living Cells
Ido’s talk at the Saturday Physics series
University of Illinois (October 2009).
Quantitative Adventures in Gene Regulation
Ido’s talk at the Workshop on Stochasticity in Biochemical Reaction Networks, Banff International Research Station (September 2013).
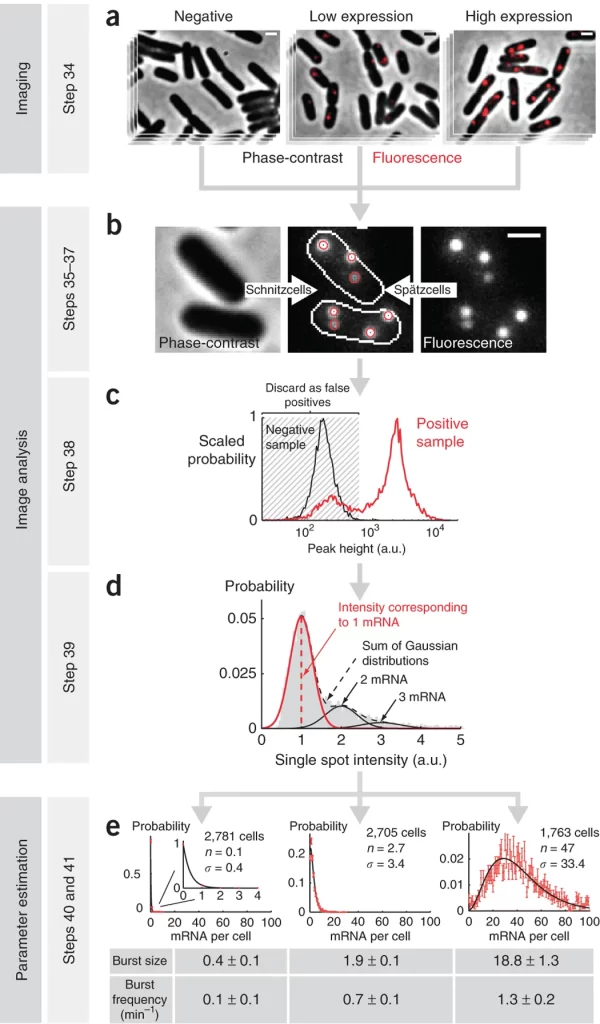
Spätzcells
Spätzcells, also known as Spaetzcells, is a MATLAB program created to analyze fluorescence images obtained from single-molecule fluorescent in situ hybridization (smFISH) experiments in E. coli.
For citations and further information, please see Skinner et al., Nature Protocols 2013 (Paper webpage; PDF).
Download
Spätzcells can be downloaded using this link (approx. 310 MB), which contains the following files:
- Codes.zip: Folder with Spätzcells code and analysis scripts.
- Example.zip: Folder containing fluorescence images and segmentation masks to run the Spätzcells tutorial.
- spatzcells_documentation.pdf: Instruction manual and tutorial.
Requirements
MATLAB (The MathWorks), as well as the MATLAB Image Processing and Optimization toolboxes, are required to run this software. Spätzcells was developed using MATLAB versions 7.10 (R2010a) and 7.13 (2011b) (both 32 and 64 bits).
